Only until a reader asked me about the mechanics of how to do a Roth IRA conversion did I realize Roth IRA “conversion” is such a misnomer. It doesn’t actually convert your IRA.
I drew some graphical illustrations to show what happens when you convert an IRA. Please note these only show the mechanics. Whether you should convert, when you should convert, and how much you should convert are out of scope in this discussion.
Container vs Content
If you have an IRA you know an IRA is just a container. You hold content inside the IRA container — stocks, bonds, mutual funds, ETFs, CDs, etc. The label on the container says what type of IRA it is: traditional IRA, Roth IRA, SEP IRA, or SIMPLE IRA. The assets in the IRA do not know or care what label is on the container. Here we have three objects in a box with a label on the box:
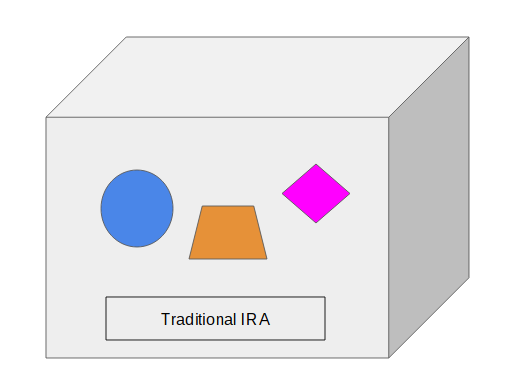
Change Label vs Move Content
Converting an IRA only goes one way: from traditional, SEP, or SIMPLE IRA to Roth IRA. When you convert the IRA, you do NOT change the label on the container like this:
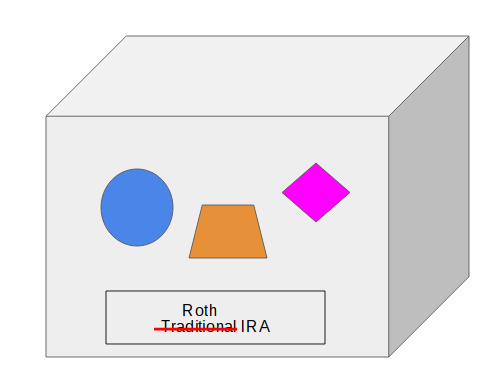
Instead, you move some or all the content in the IRA to a new or existing Roth IRA.
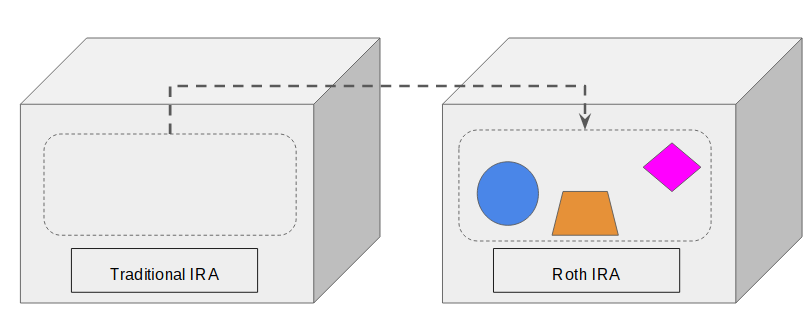
Converting a traditional IRA into a Roth IRA doesn’t really “convert” the traditional IRA. The traditional IRA is still there afterwards. It’s still the same type. It just loses some or all of its content.
You do it at the same custodian by filling out a form, either online or paper. If you don’t already have a Roth IRA with them you will open a new one. If you already have a Roth IRA with them you can choose to move the assets into the existing Roth IRA or a new one. You don’t have to sell the assets to cash unless you want to make a change in conjunction with the move. Whatever form the assets are in — stocks, bonds, mutual funds, ETFs, CDs — they can move in their current form.
Full vs Partial
If you move all the content from your traditional IRA to a Roth IRA, your traditional IRA will be empty after you are done, but it’s still there. You can put more money into it in the future.
You can also choose to move only part of the content. The traditional IRA still holds the remaining assets. It’s one more reason you don’t “convert” the IRA itself. Only the assets move.
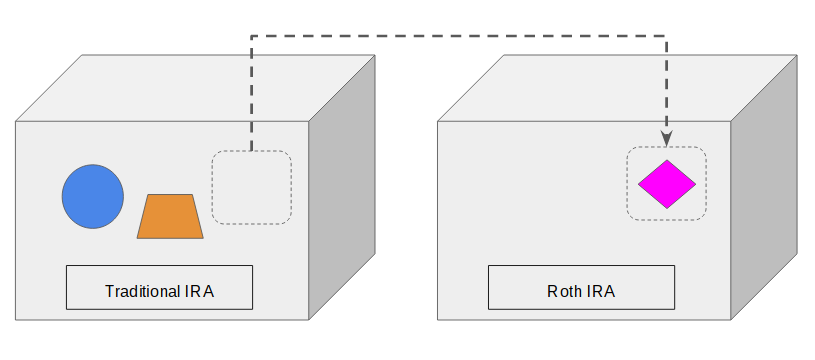
Learn the Nuts and Bolts
I put everything I use to manage my money in a book. My Financial Toolbox guides you to a clear course of action.
James says
Usually like your articles but I think this one should have never seen the light of day!
Roth move doesn’t have the same ring to it. In the end it is just a label and the assets are being converted from being in a trad ira to being in a roth ira.
m says
I agree with this comment. I was very interested until I realized this article wasn’t really saying anything at all.
msf says
Got to agree with James here. Perhaps you’d prefer “recharacterize” instead of “convert” for a Roth conversion? The investor is changing the character of the assets – from being inside one box to being inside another. Then the term for “unconverting” a Roth would become “re-recharacterization” rather than merely “recharacterization”?
Also, when you say that the empty traditional IRA will still be there, I’m guessing you mean the IRA account (yes, redundant) with the custodian, not an IRA in an abstract sense. It turns out that the IRA account won’t always be there – the custodian will shut it down completely after some waiting period. So at some point in the future, you may not be able to put more money in. You’ll have to create a new traditional IRA box.
Harry Sit says
The assets are just assets. Their characteristics are stocks, bonds, CDs, mutual funds, ETFs, and the associated risk and returns. If you ask the asset manager, iShares, for example, they don’t care or know which accounts you assets are held in. They just manage the assets. The assets stay the same in the “conversion” process. They are merely moved, not turned into something else.
I never hear people say “convert my Vanguard Total Stock Market Index Fund” only “convert my traditional IRA.” The conversion is perceived to be operated on the account, the container, not on the assets, the content, but it’s not true.
Only the accounts determine how much you can put in and what happens when you pull money out or move from one account to another. Each account has a number. The account type of each numbered account does not change/convert with the “conversion.” Even when the empty account gets closed it’s still of the original type. Its characteristic doesn’t change.
KD says
Harry, I like these kind of articles. They seem simplistic but actually bring to light nuances that are seldom noticed. Would a phrasing like asset transfer from traditional IRA to a Roth IRA be better? Bad phrasing happening/germinating from PF blogs like Backdoor Roth and Mega backdoor Roth seem to forget that labeling exercise in itself needs a thorough thought. Catchy names are easy, conveying correct meaning is hard.
Harry Sit says
Asset transfer or move or something of that nature would be a better description. You have a wooden box and a metal box, each holding some balls. When you dump some or all the balls in the wooden box into the metal box, you are not converting the wooden box into a metal box. Even when you discard the wooden box because it’s now empty and you don’t need it any more, it’s still a wooden box. It doesn’t magically turn into metal.
James says
But one could see the process as really involving a) splitting off the assets into a new tira then b) converting that tira to a roth and c) combining that with an exisiting roth if you choose or leaving it as a new roth.
That way there is an ira conversion and the old one remains as well since the assets to be included in the conversion are split off.
The trustee just simplifies the process so it looks like the assets just move from one to another.
Harry Sit says
We are talking about what actually happens in real life here, not what one could see it as. The trustee doesn’t make it “look like” the assets just move from one to another. They actually move the assets from one account to another.
The official wording in the federal regulations 26 CFR 1.408A-4 is “convert an amount in a traditional IRA to an amount in a Roth IRA.” Note the “amount” is being converted, not the IRA. The regs give three methods for doing it. The most common method is the third one:
“(3) An amount in a traditional IRA is transferred to a Roth IRA maintained by the same trustee.”
It goes on to say that if the trustee re-labels the account, it’s treated as moving the assets.
“For purposes of sections 408 and 408A, redesignating a traditional IRA as a Roth IRA is treated as a transfer of the entire account balance from a traditional IRA to a Roth IRA.”
https://www.law.cornell.edu/cfr/text/26/1.408A-4
Because it’s treated as moving the assets anyway, in the real world the trustee just moves the assets, which is the officially sanctioned procedure. The account is not split, converted, or merged.
serbeer says
Running low on ideas Harry? 🙂
Harry Sit says
More ideas are always welcome. If you have done it multiple times year after year it may not be news to you. To someone who have never done it, when they hear about converting an IRA, they naturally get the impression that the IRA itself gets converted, not just the assets in it get moved. When you throw in rollover, transfer, and recharacterize, it gets more confusing. More diagrams are coming.
MoneyOCD says
Surprisingly, I see no value in that article 🙁 it does not matter what you name what, what matter is that pretax dollar became after tax dollar – same dollars (!!!!) not a conversion? fine, we can name it “tax magic” 🙂
Harry Sit says
Don’t confuse the container with its content. After you convert your IRA, when you contribute more money to the same account, are you contributing to traditional or to Roth? Traditional? Didn’t you just convert your IRA so it became Roth?